Threads fitting data#
From the threads documentation regarding fitting input mutations to the inferred ARG:
Threads does not by default guarantee that input mutations correspond to monophyletic clades in the inferred ARG. However, this can be achieved by providing the
--fit_to_dataflag. When threads infer is run with the--fit_to_dataflag, the threading instructions are amended post-hoc such that each input mutation is represented by a clade in a marginal coalescence tree. As part of this process, Threads estimates the age of each mutation and then amends the local ARG topology.
However, the time required to fit the data can be significant, especially for large datasets.
Another consideration is that when converting the ARG to a tree sequence, I can’t
use the --add_mutations if --fit_to_data is not used: this is why the
nf-treeseq pipeline currently models
the two distinct options with --threads_fit_to_data parameter.
import tszip
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from tskitetude import get_project_dir
Load threads tree sequences#
Let’s open the simplest results I have, with and without fitting the data, and try to understand the differences:
ts_threads = tszip.load(get_project_dir() / "results-threads/toInfer/threads/ts300I25k.1.trees.tsz")
ts_threads_fit = tszip.load(get_project_dir() / "results-threads/toInfer-fit/threads/ts300I25k.1.trees.tsz")
Let’s dump the two tree sequences:
ts_threads
|
|
|
|---|---|
| Trees | 22 554 |
| Sequence Length | 99 988 756 |
| Time Units | unknown |
| Sample Nodes | 3 200 |
| Total Size | 145.5 MiB |
| Metadata |
dictchromosome: 1offset: 6714 |
| Table | Rows | Size | Has Metadata |
|---|---|---|---|
| Edges | 3 446 739 | 105.2 MiB | |
| Individuals | 1 600 | 80.8 KiB | ✅ |
| Migrations | 0 | 8 Bytes | |
| Mutations | 0 | 16 Bytes | |
| Nodes | 520 417 | 13.9 MiB | |
| Populations | 8 | 271 Bytes | ✅ |
| Provenances | 1 | 385 Bytes | |
| Sites | 0 | 16 Bytes |
| Provenance Timestamp | Software Name | Version | Command | Full record |
|---|---|---|---|---|
| 15 December, 2025 at 02:21:32 PM | threads | 0.2.1 | Unknown |
Detailsdict
software:
dictname: threadsversion: 0.2.1
parameters:
dictinput_file: ts300I25k.1.biallelic.vcf.gzmetadata_added: True populations_added: 8 individuals_added: 1600 timestamp: 2025-12- 15T14:21:32.293625+00:00 description: Tree sequence annotated with population and individual metadata |
ts_threads_fit
|
|
|
|---|---|
| Trees | 24 857 |
| Sequence Length | 99 988 756 |
| Time Units | unknown |
| Sample Nodes | 3 200 |
| Total Size | 354.1 MiB |
| Metadata |
dictchromosome: 1offset: 6714 |
| Table | Rows | Size | Has Metadata |
|---|---|---|---|
| Edges | 8 329 337 | 254.2 MiB | |
| Individuals | 1 600 | 80.8 KiB | ✅ |
| Migrations | 0 | 8 Bytes | |
| Mutations | 24 924 | 900.6 KiB | |
| Nodes | 1 304 983 | 34.8 MiB | |
| Populations | 8 | 271 Bytes | ✅ |
| Provenances | 1 | 385 Bytes | |
| Sites | 24 924 | 608.5 KiB |
| Provenance Timestamp | Software Name | Version | Command | Full record |
|---|---|---|---|---|
| 17 December, 2025 at 08:29:37 PM | threads | 0.2.1 | Unknown |
Detailsdict
software:
dictname: threadsversion: 0.2.1
parameters:
dictinput_file: ts300I25k.1.biallelic.vcf.gzmetadata_added: True populations_added: 8 individuals_added: 1600 timestamp: 2025-12- 17T20:29:37.025492+00:00 description: Tree sequence annotated with population and individual metadata |
The fitted version seems bigger than the normal one. Let’s open also
the tsinfer version of treeseq:
ts_infer = tszip.load(get_project_dir() / "results-reference/toInfer/tsinfer/ts300I25k.1.trees.tsz")
ts_infer
|
|
|
|---|---|
| Trees | 22 569 |
| Sequence Length | 1e+08 |
| Time Units | generations |
| Sample Nodes | 3 200 |
| Total Size | 123.5 MiB |
| Metadata |
dict |
| Table | Rows | Size | Has Metadata |
|---|---|---|---|
| Edges | 1 224 660 | 37.4 MiB | |
| Individuals | 1 600 | 82.2 KiB | ✅ |
| Migrations | 0 | 8 Bytes | |
| Mutations | 851 484 | 68.8 MiB | ✅ |
| Nodes | 80 645 | 6.7 MiB | ✅ |
| Populations | 8 | 185 Bytes | ✅ |
| Provenances | 4 | 2.8 KiB | |
| Sites | 24 947 | 1.2 MiB | ✅ |
| Provenance Timestamp | Software Name | Version | Command | Full record |
|---|---|---|---|---|
| 05 December, 2025 at 09:41:56 AM | tsdate | 0.2.4 | variational_gamma |
Detailsdictschema_version: 1.0.0
software:
dictname: tsdateversion: 0.2.4
parameters:
dictmutation_rate: 5.87e-09recombination_rate: None time_units: None progress: None population_size: None eps: 1e-10 max_iterations: 25 max_shape: 1000 rescaling_intervals: 1000 rescaling_iterations: 5 match_segregating_sites: False regularise_roots: True singletons_phased: True command: variational_gamma
environment:
dict
os:
dictsystem: Linuxnode: node1 release: 5.15.0-58-generic version: #64-Ubuntu SMP Thu Jan 5 11:43:13 UTC 2023 machine: x86_64
python:
dictimplementation: CPythonversion: 3.12.12
libraries:
dict
tskit:
dictversion: 1.0.0b3
resources:
dictelapsed_time: 108.51660299301147user_time: 571.01 sys_time: 14.32 max_memory: 1730285568 |
| 05 December, 2025 at 09:40:07 AM | tsdate | 0.2.4 | preprocess_ts |
Detailsdictschema_version: 1.0.0
software:
dictname: tsdateversion: 0.2.4
parameters:
dictminimum_gap: 1000000erase_flanks: True split_disjoint: True filter_populations: False filter_individuals: False filter_sites: False
delete_intervals:
listlist06713.0 list99995468.0100000000.0 command: preprocess_ts
environment:
dict
os:
dictsystem: Linuxnode: node1 release: 5.15.0-58-generic version: #64-Ubuntu SMP Thu Jan 5 11:43:13 UTC 2023 machine: x86_64
python:
dictimplementation: CPythonversion: 3.12.12
libraries:
dict
tskit:
dictversion: 1.0.0b3
resources:
dictelapsed_time: 6.797327280044556user_time: 462.99 sys_time: 13.57 max_memory: 1596768256 |
| 05 December, 2025 at 09:40:00 AM | tskit | 1.0.0b3 | simplify |
Detailsdictschema_version: 1.0.0
software:
dictname: tskitversion: 1.0.0b3
parameters:
dictcommand: simplifyTODO: add simplify parameters
environment:
dict
os:
dictsystem: Linuxnode: node1 release: 5.15.0-58-generic version: #64-Ubuntu SMP Thu Jan 5 11:43:13 UTC 2023 machine: x86_64
python:
dictimplementation: CPythonversion: 3.12.12
libraries:
dict
kastore:
dictversion: 2.1.1 |
| 05 December, 2025 at 09:39:55 AM | tsinfer | 0.5.0 | infer |
Detailsdictschema_version: 1.0.0
software:
dictname: tsinferversion: 0.5.0
parameters:
dictmismatch_ratio: Nonepath_compression: True precision: None post_process: None command: infer
environment:
dict
libraries:
dict
zarr:
dictversion: 2.18.3
numcodecs:
dictversion: 0.13.1
lmdb:
dictversion: 1.7.5
tskit:
dictversion: 1.0.0b3
os:
dictsystem: Linuxnode: node1 release: 5.15.0-58-generic version: #64-Ubuntu SMP Thu Jan 5 11:43:13 UTC 2023 machine: x86_64
python:
dictimplementation: CPython
version:
list312 12
resources:
dictelapsed_time: 143.15887202322483user_time: 303.99 sys_time: 10.53 max_memory: 1596768256 |
Simplify threads tree sequences#
Are threads file simplified? try to simplify them:
ts_threads_simplified = ts_threads.simplify()
ts_threads_fit_simplified = ts_threads_fit.simplify()
let’s collect those information in a DataFrame for easier comparison:
ts_list = [
ts_threads,
ts_threads_simplified,
ts_threads_fit,
ts_threads_fit_simplified,
ts_infer
]
comparison_data = {
'num_trees': [getattr(ts, 'num_trees') for ts in ts_list],
'num_edges': [getattr(ts, 'num_edges') for ts in ts_list],
'num_mutations': [getattr(ts, 'num_mutations') for ts in ts_list],
'num_nodes': [getattr(ts, 'num_nodes') for ts in ts_list],
'num_sites': [getattr(ts, 'num_sites') for ts in ts_list]
}
df_comparison = pd.DataFrame(comparison_data, index=[
'threads',
'threads-simplified',
'threads-fit',
'threads-fit-simplified',
'tsinfer'])
df_comparison.transpose()
| threads | threads-simplified | threads-fit | threads-fit-simplified | tsinfer | |
|---|---|---|---|---|---|
| num_trees | 22554 | 22554 | 24857 | 24857 | 22569 |
| num_edges | 3446739 | 1780159 | 8329337 | 4239485 | 1224660 |
| num_mutations | 0 | 0 | 24924 | 24924 | 851484 |
| num_nodes | 520417 | 520417 | 1304983 | 1304983 | 80645 |
| num_sites | 0 | 0 | 24924 | 24924 | 24947 |
Plot tree sequence stats#
Let’s plot this dataframe:
# A plot for each metric in a 2x3 grid with colors for each method (log scale)
metrics = df_comparison.columns
fig, axes = plt.subplots(2, 3, figsize=(15, 8))
axes = axes.flatten()
# Define colors for each method
colors = sns.color_palette("Set1", 5)
for i, metric in enumerate(metrics):
data = df_comparison[[metric]].reset_index()
data.columns = ['method', 'value']
if metric in ['num_mutations']:
data['value'] = np.log10(data['value'] + 1)
axes[i].set_ylabel('log10(Count)')
else:
axes[i].set_ylabel('Count')
sns.barplot(data=data, x='method', y='value', hue='method', ax=axes[i], palette=colors, legend=False)
axes[i].set_title(metric)
axes[i].set_xlabel('')
axes[i].tick_params(axis='x', rotation=45)
# Hide the last subplot (since we only have 5 metrics)
axes[5].set_visible(False)
plt.tight_layout()
plt.show()
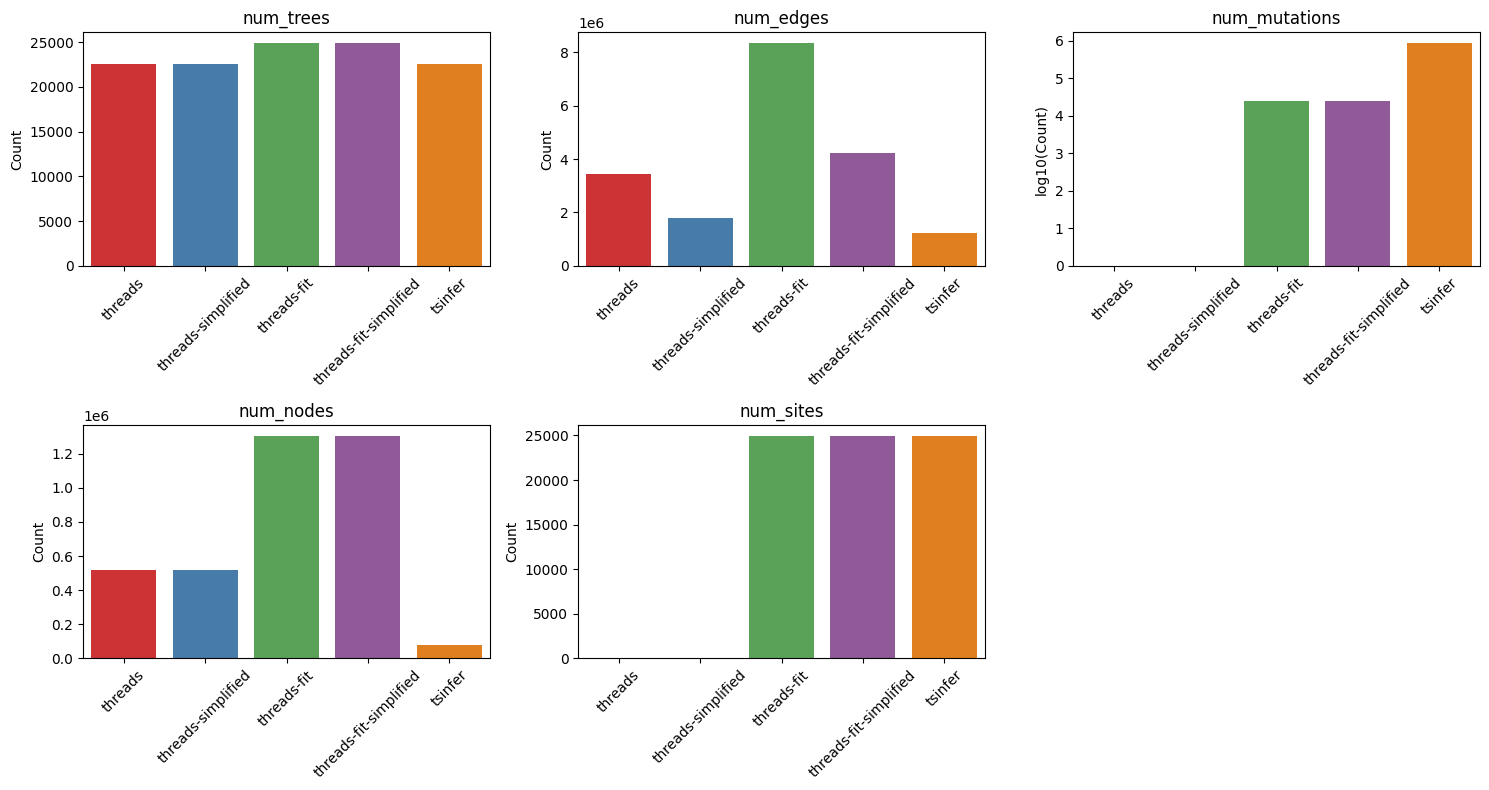
The number of trees are similar, between the two approaches (simplified or not). Number of edges differs: The unfitted version has less edges and simplified version has a number of edges similar to the tsinfer version. The fitted version and the fitted-simplified version have four and two times more edges than the tsinfer version. The number of mutations and sites are missed in the unfitted version, while the fitted version has more than half the mutations of the tsinfer version and the number of sites is similar. Lastly, the number of nodes is significantly higher in the unfitted version and more higher in the fitted version compared to the tsinfer one.
